
Developing advanced electrically active cell models is challenging. Can you capture the complexity of your stem cell-derived neuronal function with just imaging or expression data? Track the emergence of neuronal activity and watch as complex synchronous patterns emerge. Quantify network properties and classify how your cells behave.
Over minutes or months, gain unprecedented access to neuronal activity with the Maestro MEA platform. Noninvasively monitor cells in culture as they mature and establish their unique phenotype. Measure dozens of endpoints with AxIS Navigator to fully classify and characterize your model's activity.
-
Local Field Potential (LFP)>
-
Functionally optimize neural differentiation and culture conditions>
-
Identify unique network activity patterns with cell cultures from specific brain regions>
-
Assay Steps>
Neuronal cultures produce complex patterns of activity as the network matures over time. The neuronal activity can be separated into network spiking activity, characterized by the number and synchrony of detected action potentials, and the local field potential (LFP), which detects low-frequency oscillations in the network. The LFP and network activity are measured simultaneously (see left) and provide complementary information on the maturation of the network. Classical network activity measures detect activity, synchrony, and oscillations in this network of rodent cortical neurons by 14 days in culture. From the average LFP waveforms (see right), we can see that the complexity of those network events increases as the cultures mature.
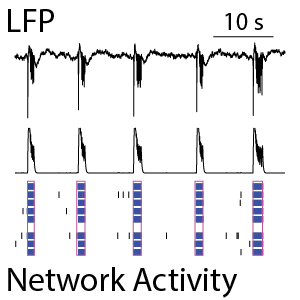
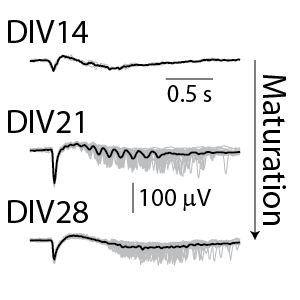
(Left) The raster plot (bottom) identified bursts of spiking detected on individual electrodes (blue) and coordinated bursts of activity across electrodes (pink) for rodent cortical neurons after 21 days in culture. The oscillations in the network activity were detected via in the population activity histogram (middle). The local field potential (LFP) signal (top) was measured simultaneously from each electrode in the well, with one example trace depicted here. The LFP events were coordinated with the network activity in the well. (Right) The detected LFP events (gray) are presented along with the average LFP across events (black) at different stages of neuronal network maturation for rodent cortical neurons. At 14 days in vitro (DIV), the LFP events are small, biphasic, and short in duration. At DIV21, the LFP reveals oscillations in the neuronal network. At 28 days in vitro, the network displayed strong initial peaks with rebound events occurring at variable delays.
Action potential firing and synaptic activity are fundamental properties of neurons in the brain. Bardy et al. [PNAS, 2015] have recently reported that Neurobasal® Medium and DMEM/F-12 support neuron survival but suppress their synaptic activities in culture. Since it is desirable for human pluripotent stem cell (hPSC)-derived neurons to have spontaneous electrical activity BrainPhys Neuronal Medium, was developed. Here the effect of BrainPhys™ and DMEM/F-12 based media on the network activity of hPSC-derived neurons during 18 week in culture was tested.
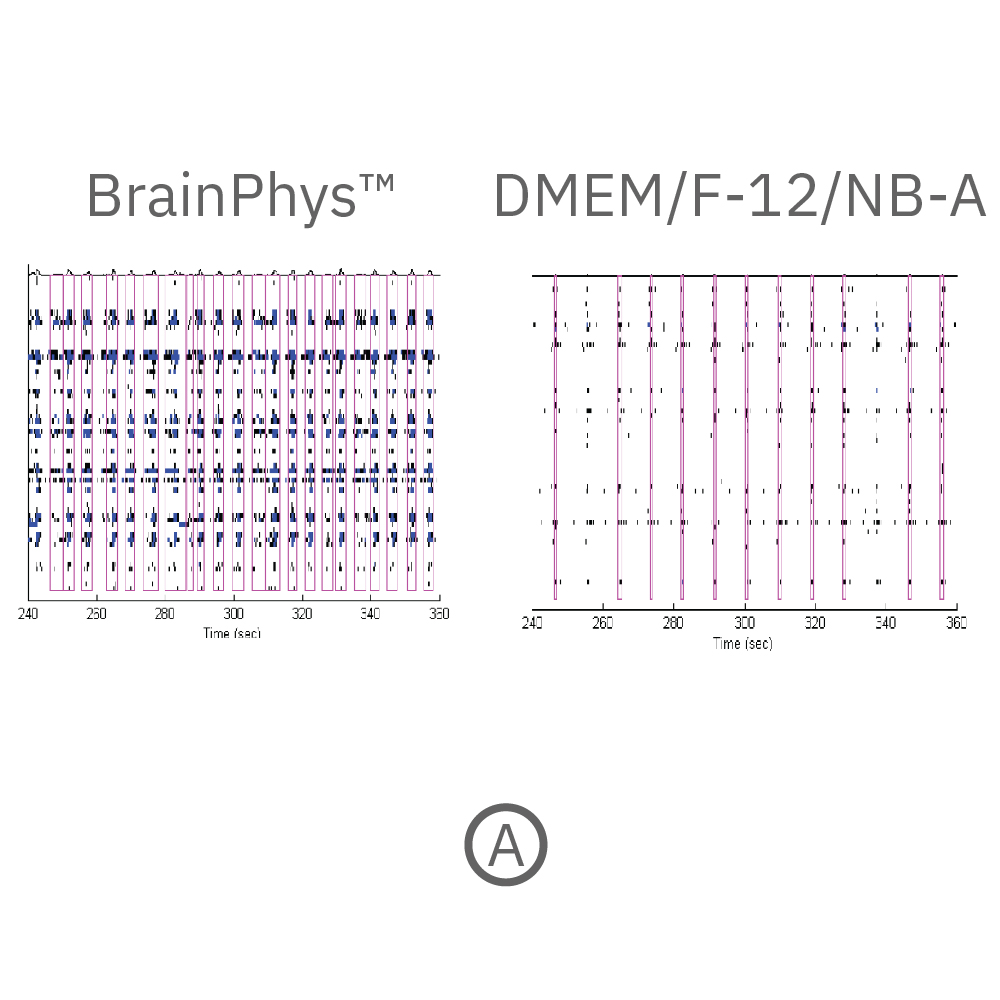
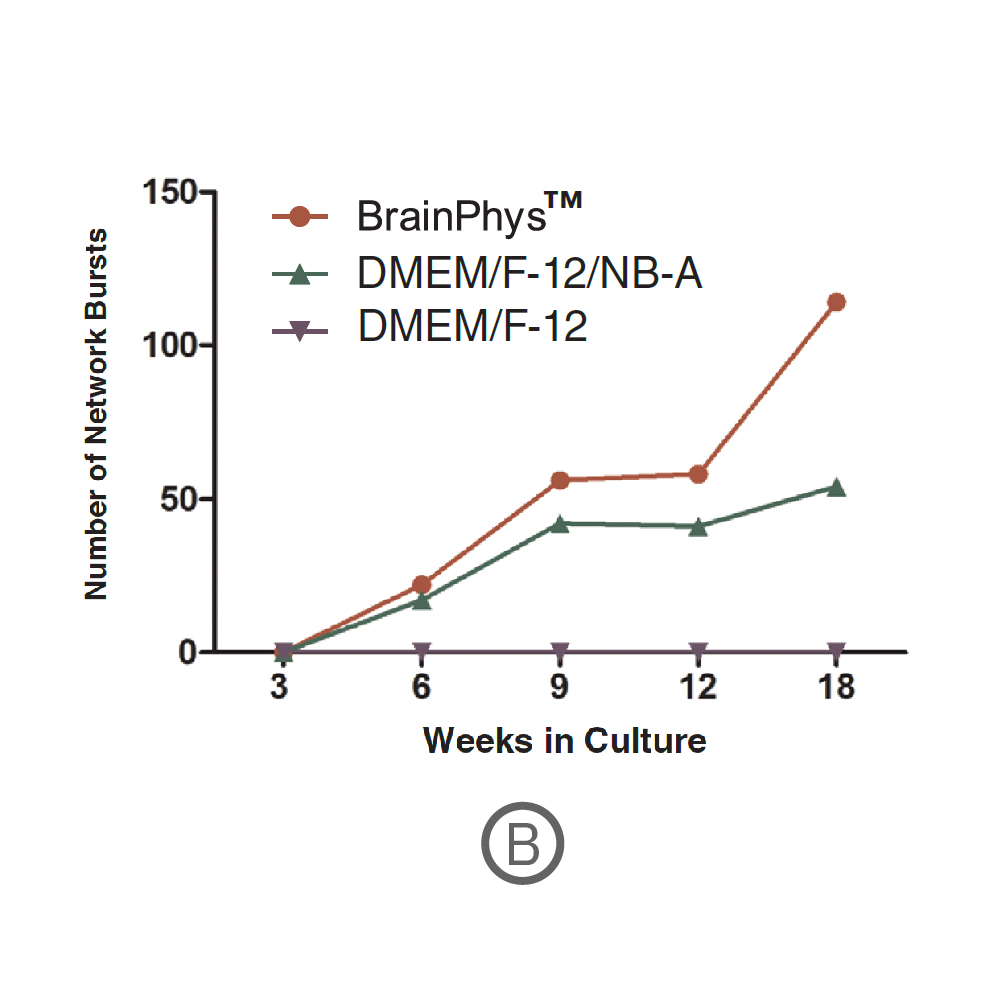
(A) Raster plots showing the firing patterns of neurons across 64 electrodes after 18 weeks in culture. Each black line represents a detected spike. Blue lines represent single channel bursts - a collection of at least 5 spikes, each separated by an inter-spike interval (ISI) of no more than 100 ms. Network bursts are marked with purple boxes and are defined as a collection of at least 10 spikes from a minimum of 25% of participating electrodes across the entire well, each separated by an ISI of no more than 100 ms. Neurons cultured in BrainPhys had increased firing compared to DMEM/F-12/NB-A. (B) Network bursts were first detected at week 6 in BrainPhys™ and DMEM/F-12/NB-A conditions. The number of network bursts increased gradually over time, suggesting that both cultures became more synchronous as they matured. After 18 weeks in culture, the number of network bursts detected in a 10-minute recording in BrainPhys™ and DMEM/F-12/NB-A were 114 and 54, respectively. Data courtesy of STEMCELL Technologies, taken from Mak et al. 2016 presented at SfN2016.
Differences in electrical activity patterns from primary murine neuronal cell cultures were compared on the Maestro MEA system. As a result of their phenotypic receptor and neuron type composition, primary neuronal cell cultures show very specific and complex activity patterns after four weeks in vitro. This complexity results from a high level of network organization in primary cultures.
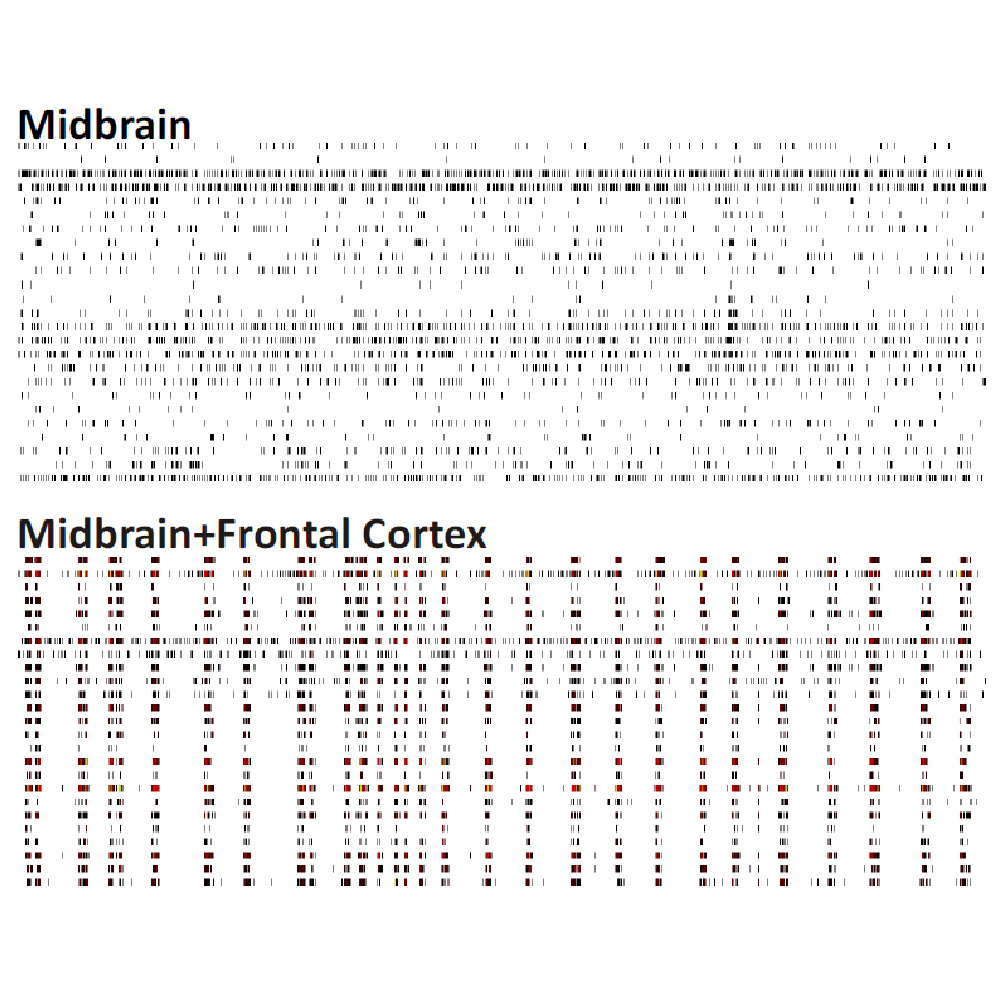
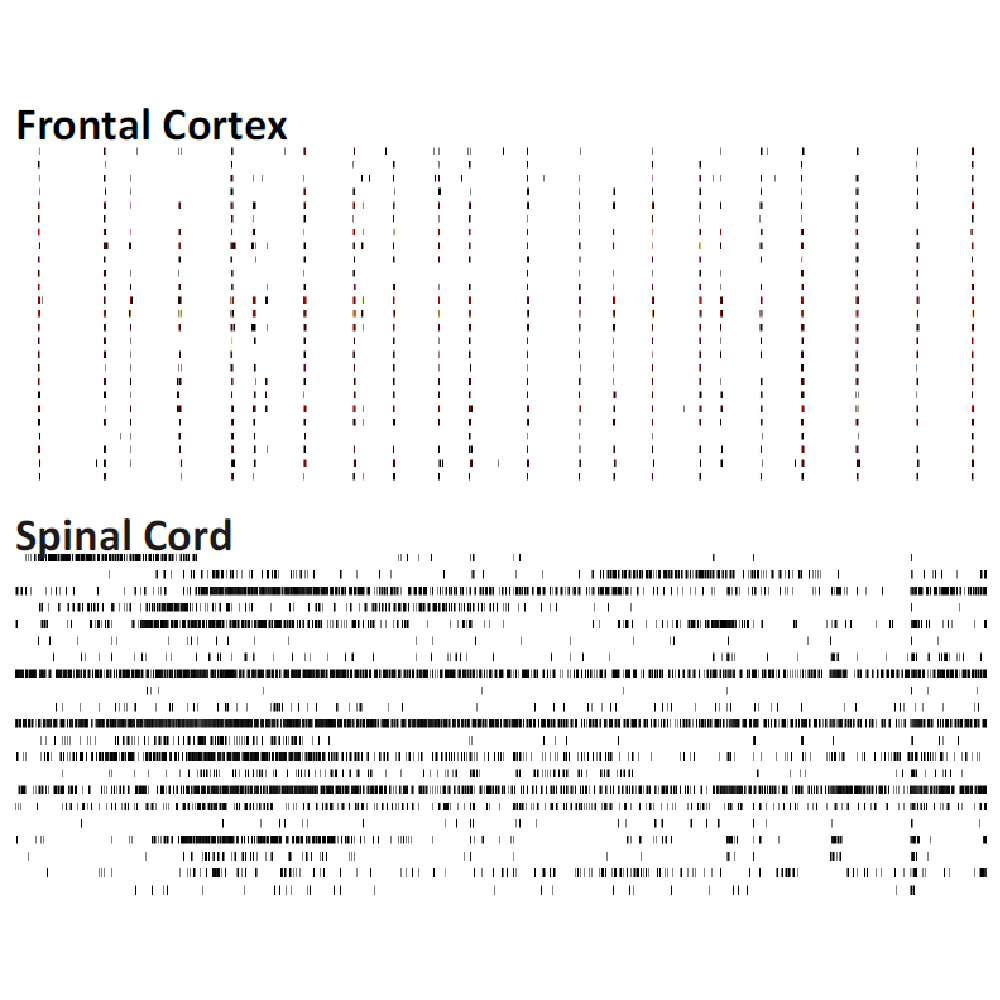
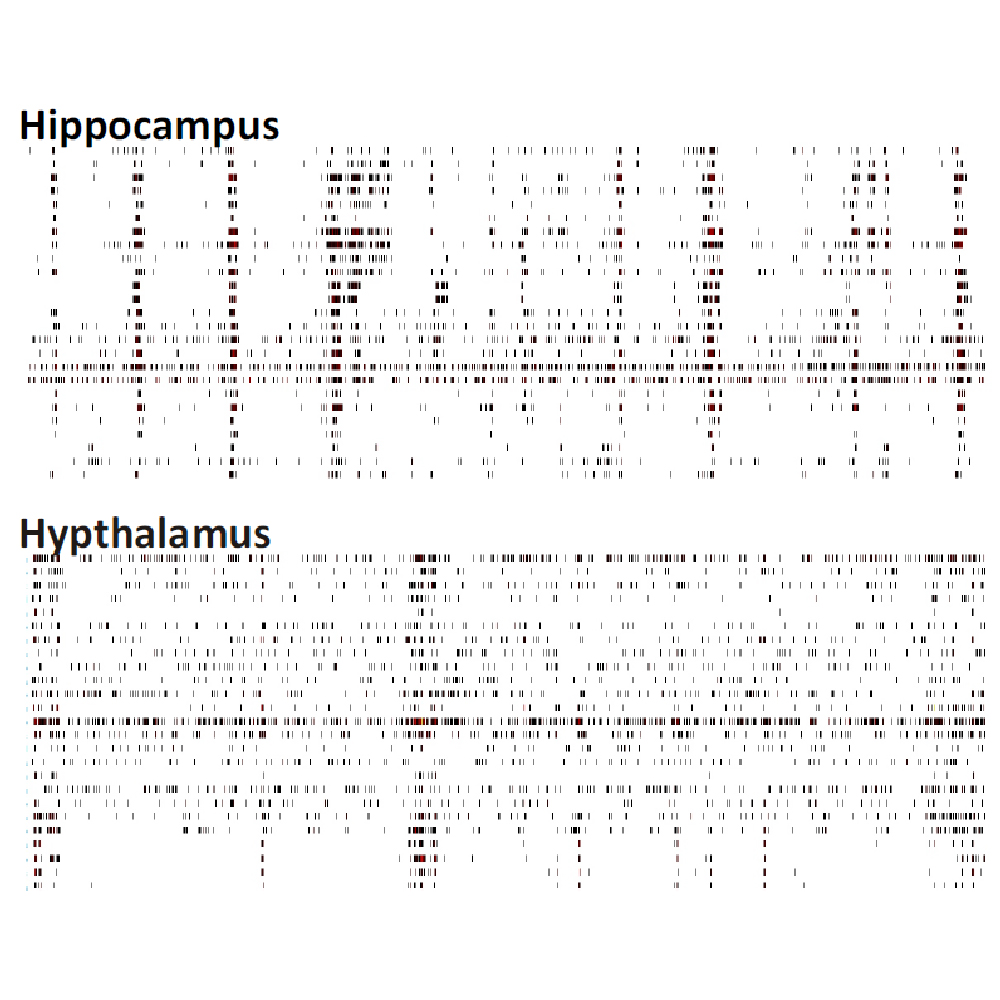

Raster plots of brain region-specific primary cell cultures derived from embryonic murine tissue of the frontal cortex, spinal cord (with dorsal root ganglia), hippocampus, and midbrain (co-cultured with frontal cortex) were compared. Plotted are 60 seconds of 25 neurons of spontaneous network activity at 28 days in vitro. Spontaneous activity plotted for 60 seconds shows distinguishable patterns based on the brain region of origin. Data courtesy of Neuroproof GMBH, taken from Voss et al. 2014 presented at SfN2014.
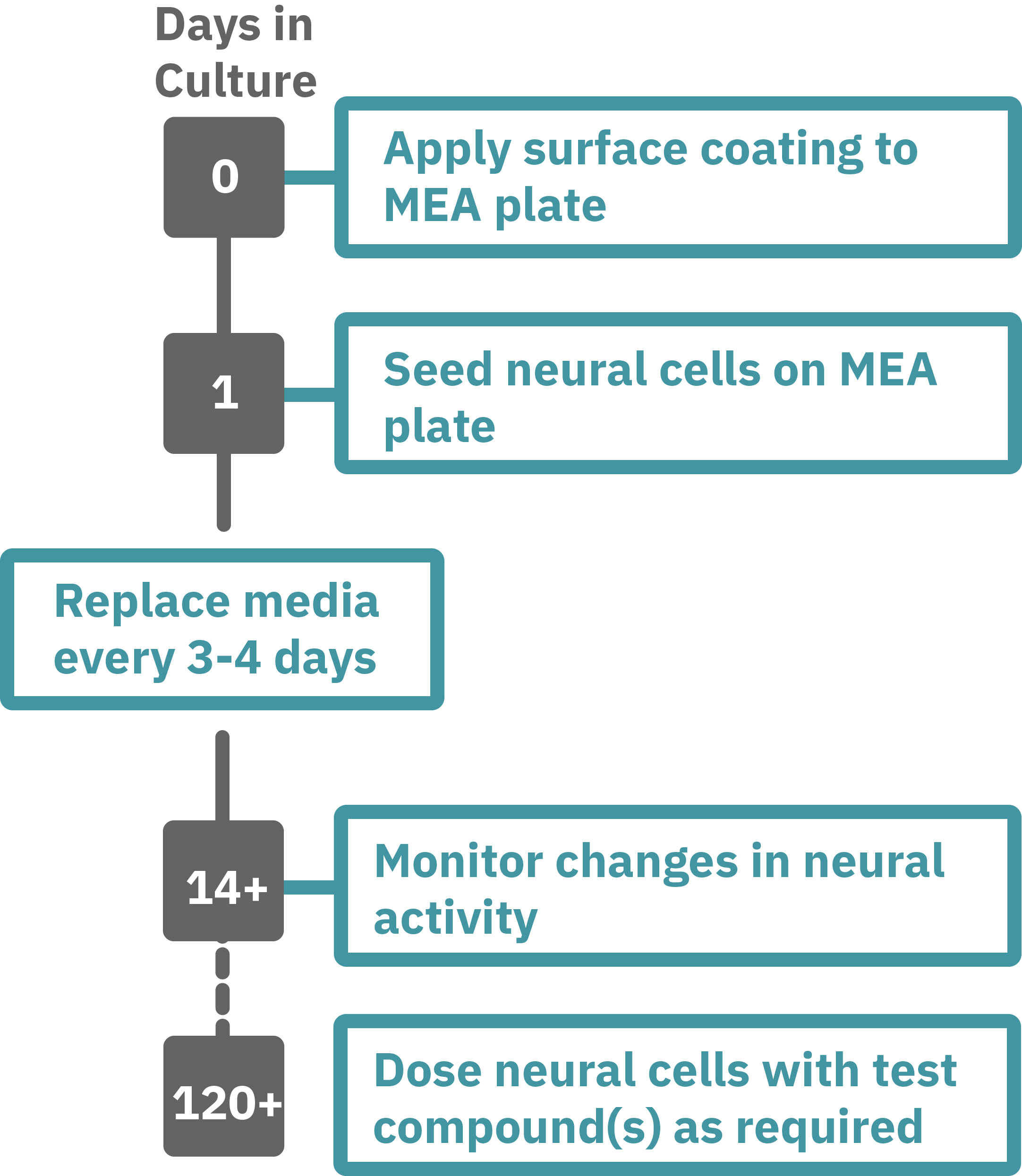
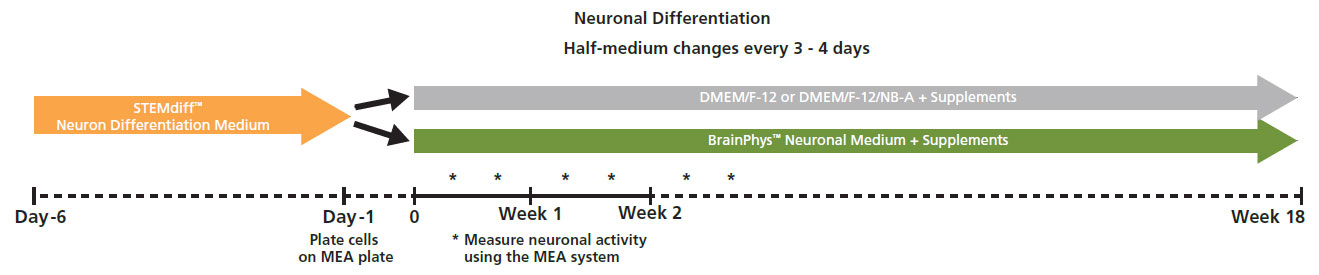
Protocol taken from Mak et al. 2016 presented at SfN2016. Neural progenitor cells derived from hPSCs (XCL1-NPC) were cultured in STEMdiff™ Neuron Differentiation Medium on poly-L-ornithine (PLO)/laminin-coated 6-well plate for 5 days. On day 5, neural progenitor cells were dissociated and single cells were re-plated onto a PLO/laminin-coated CytoView MEA plates at 30,000 cells/cm2 in STEMdiff™ Neuron Differentiation Medium. After one day, half of the medium was replaced with differentiation media (BrainPhys™ Neuronal Medium + supplements: 1% N2 Supplement-A, 2% NeuroCult™ SM1 Neuronal Supplement, 20 ng/mL GDNF, 20 ng/mL BDNF, 1 mM db-cAMP and 200 nM Ascorbic Acid). Half-medium changes were performed every 3 - 4 days throughout the culture period. Spontaneous neuronal activity was acquired at 37°C under a 5% CO₂ atmosphere using the Maestro MEA system. A 15-minute recording was taken twice a week and analyzed with AxIS Navigator Neural Module software.








