Developing advanced electrically active cell models is challenging. Can you capture the complexity of your stem cell-derived cardiac function with just imaging or expression data? Track the emergence of cardiac activity and watch as the syncytium forms and matures. Quantify beat properties and classify how your cells behave. The Maestro Pro and Edge MEA systems provides you with the cell activity information you've been missing. Over minutes or months, gain unprecedented access to cardiac activity with the Maestro MEA platform. Noninvasively monitor cells in culture as they mature and establish their unique phenotype. Measure a dozen endpoints with AxIS Navigator to fully classify and characterize your model's activity.
Human pluripotent stem cell-derived cardiomyocytes (hPSC-CMs) are an attractive model for studying heritable and drug-acquired arrhythmias. The ability to efficiently produce multiple patient-derived hPSC-CMs and easily track their performance would facilitate studies of the role of genetic diversity on arrhythmogenesis. In pursuit of this objective, hPSC differentiation into cardiomyocytes using the STEMdiff™ Cardiomyocyte Differentiation and Maintenance Kits (STEMCELL Technologies) was tracked each day using the Maestro MEA system.
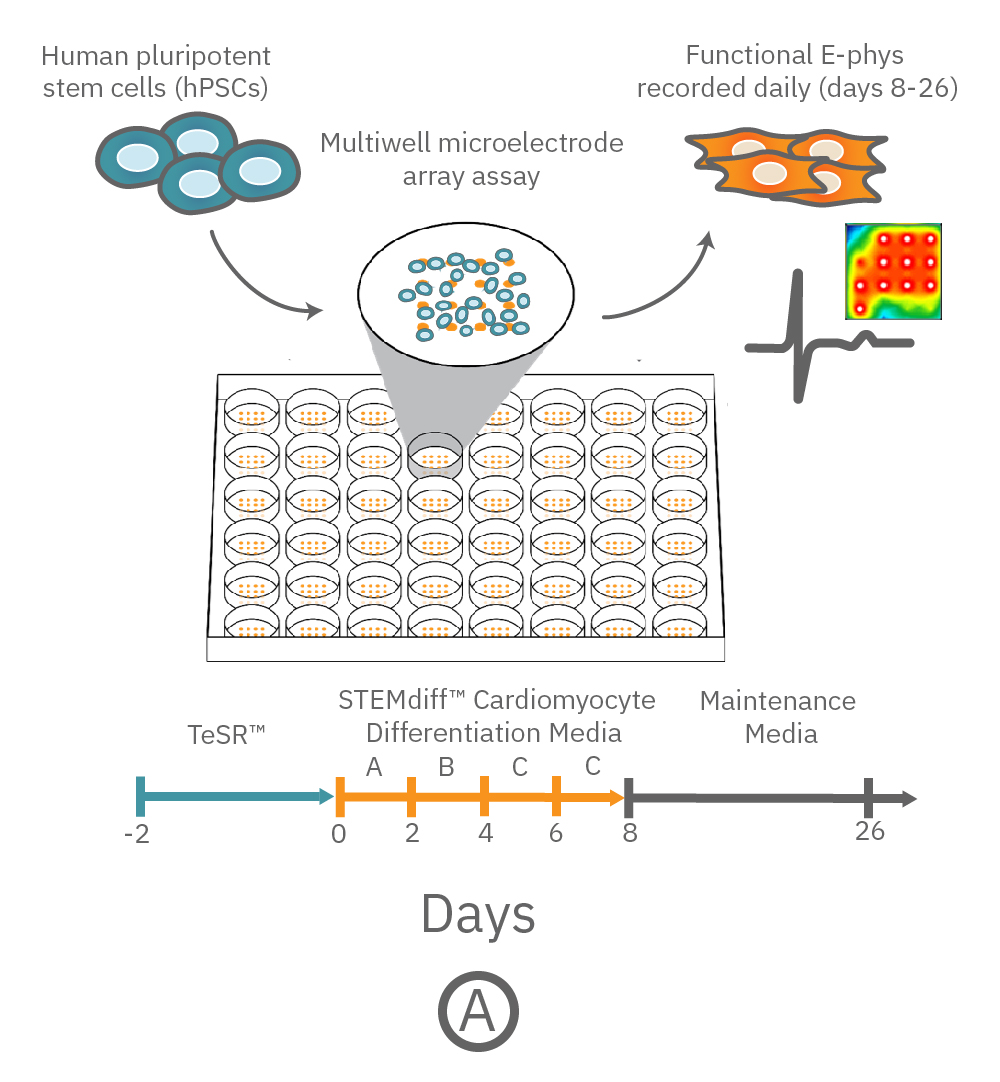
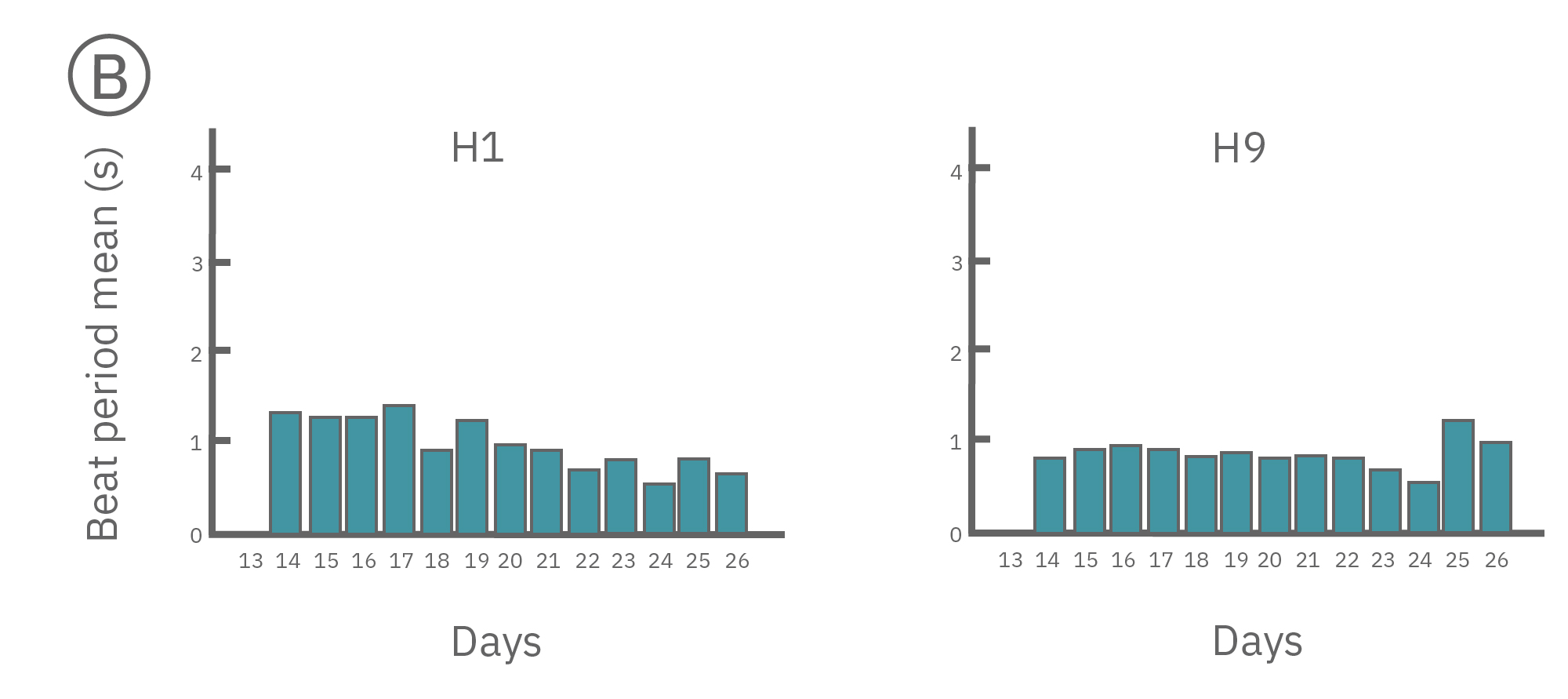
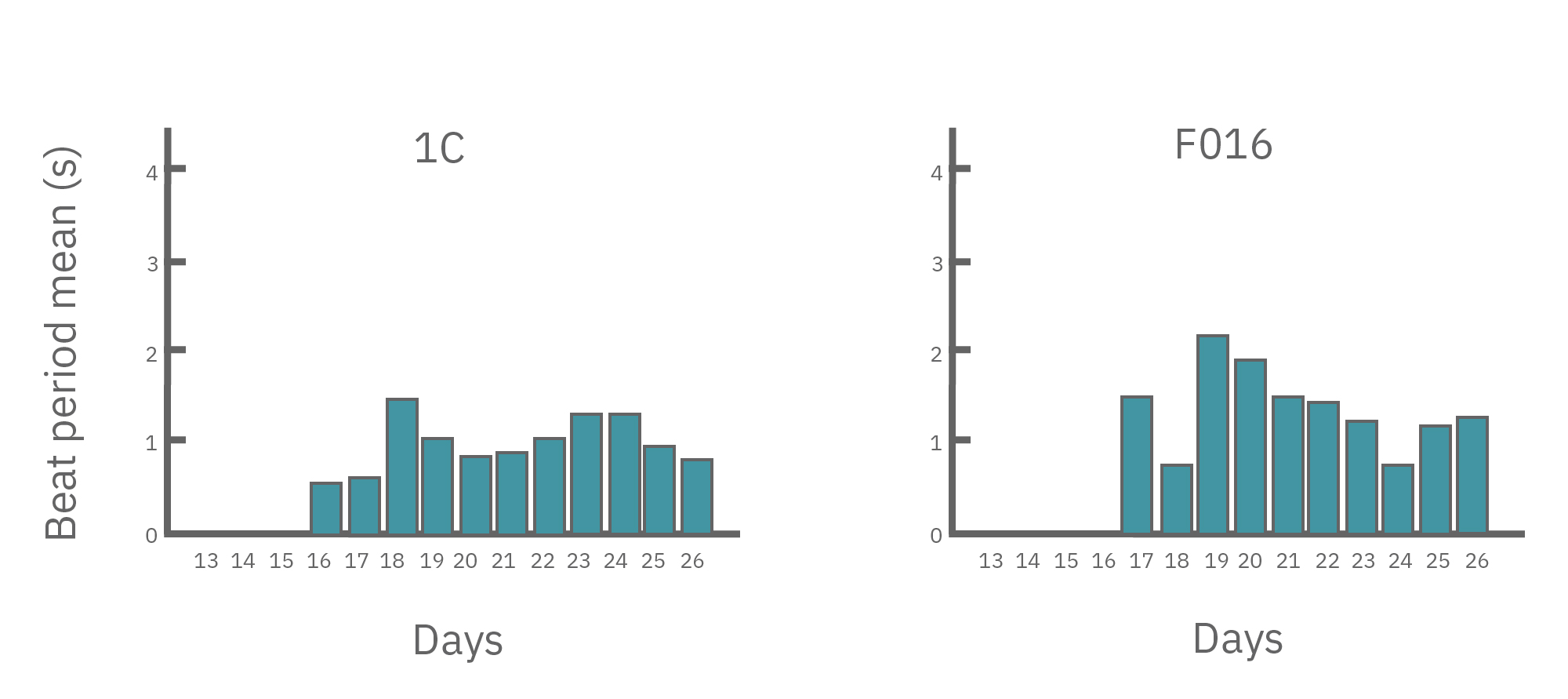
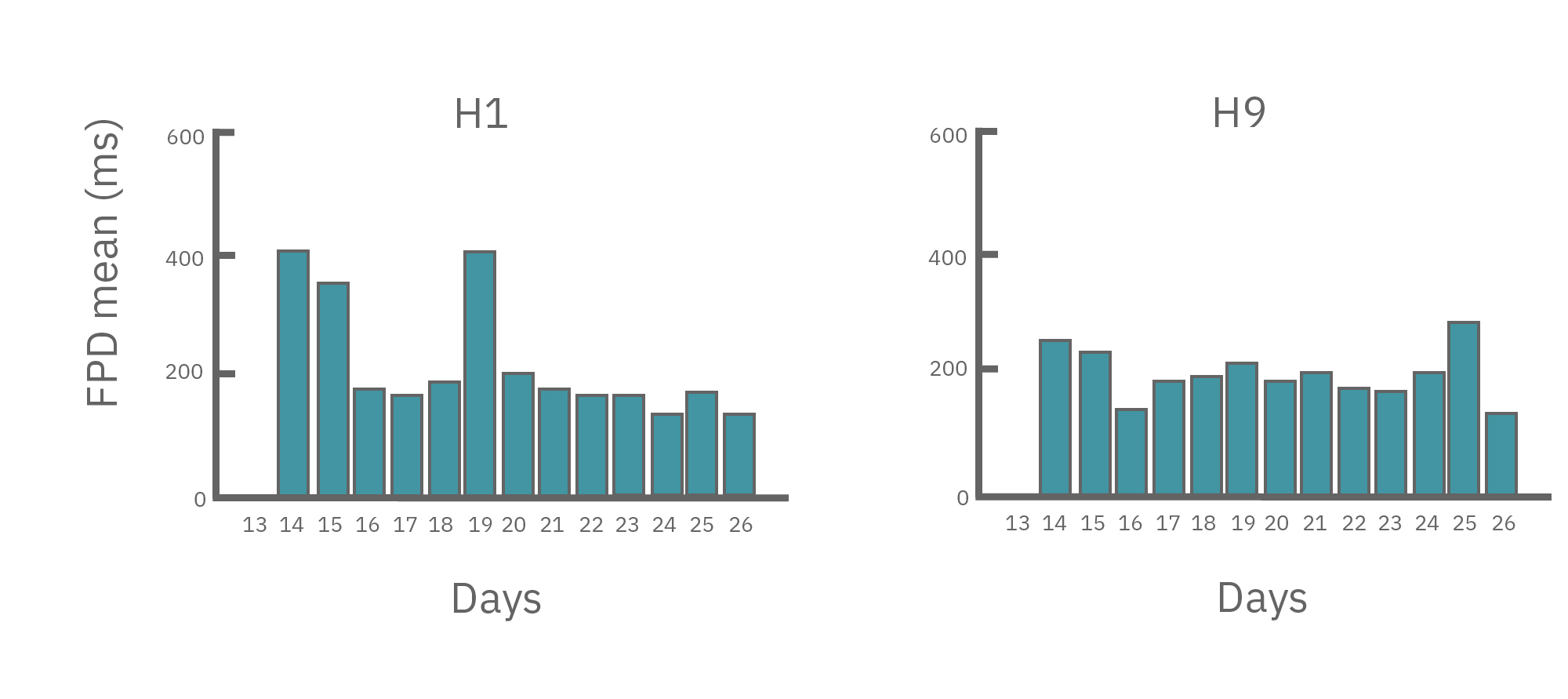
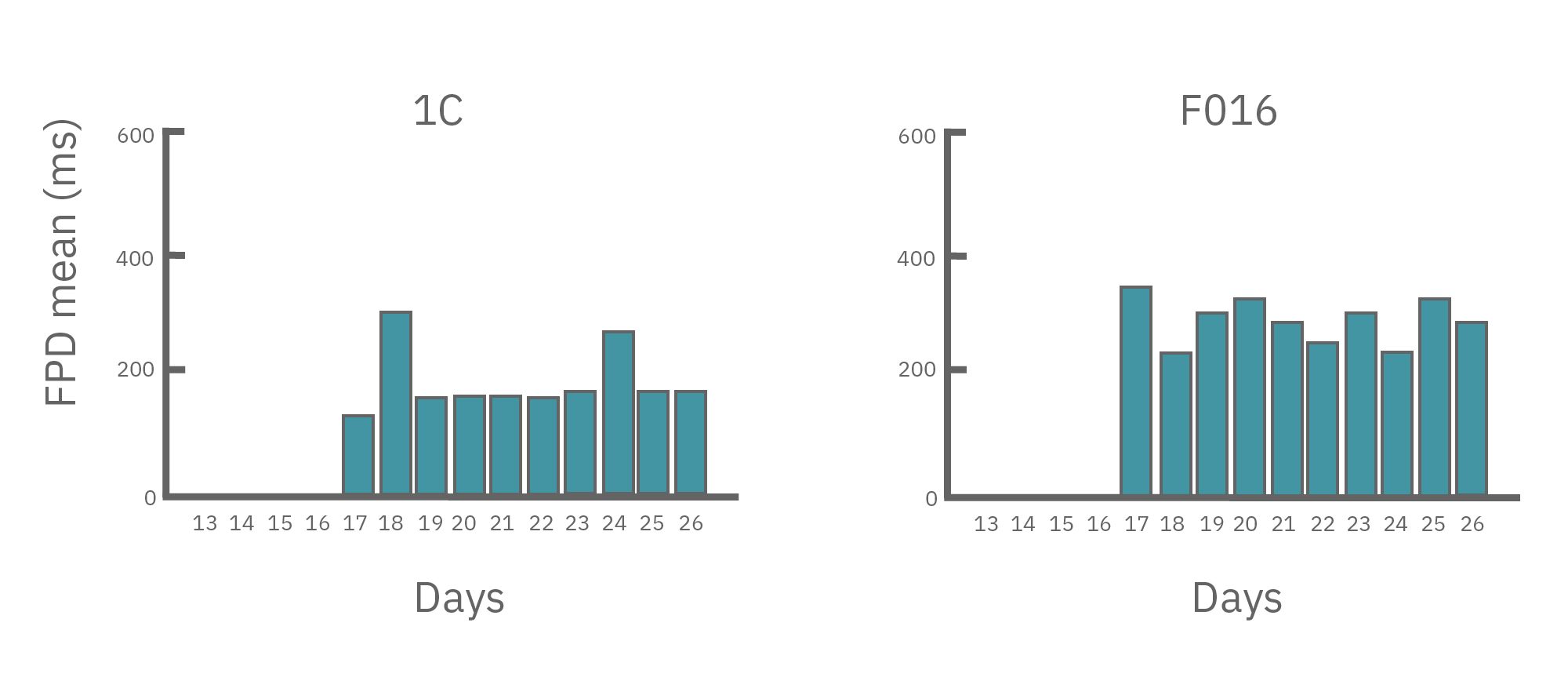
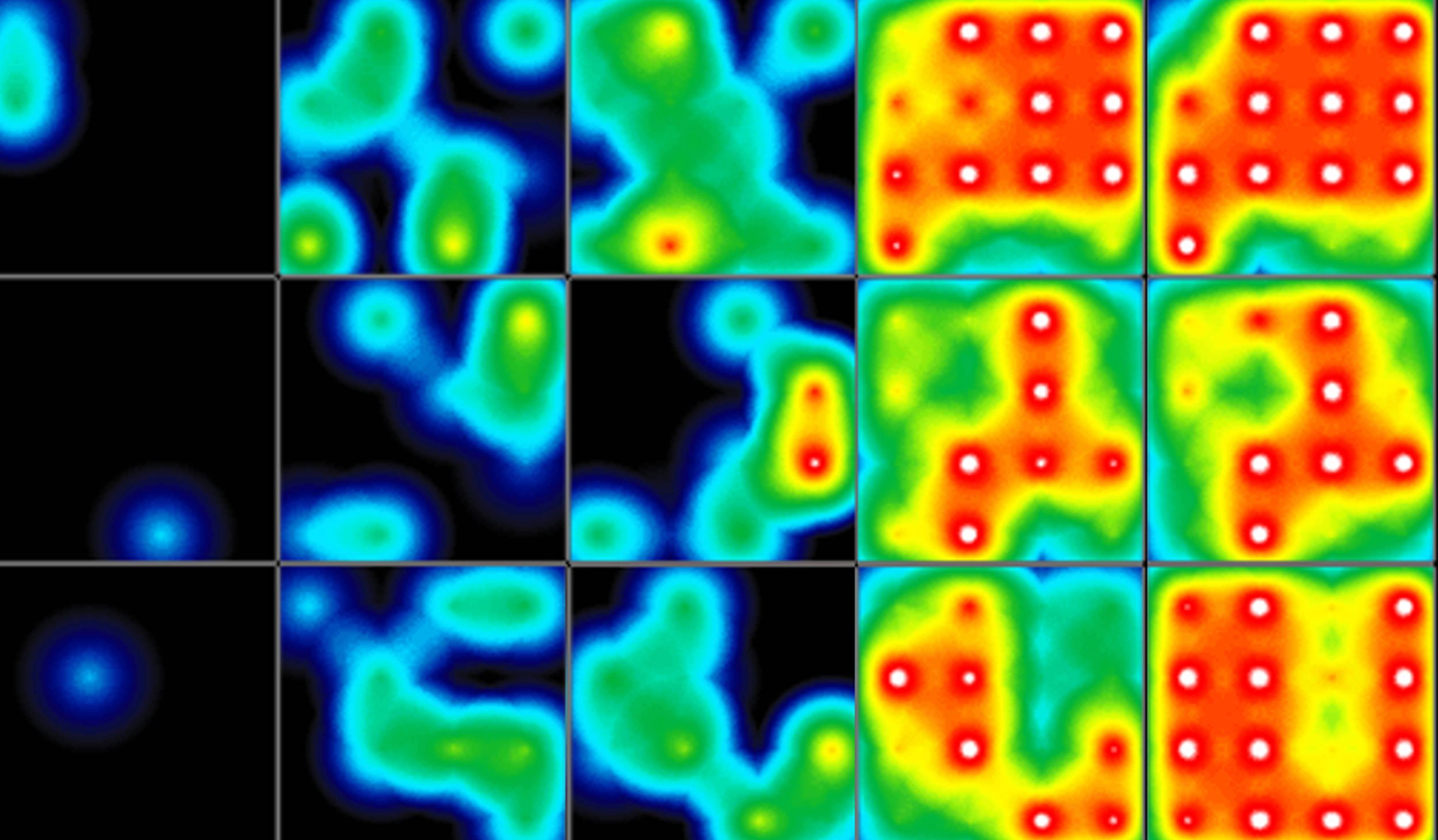
(A) Protocol used to differentiate hPSCs into cardiomyocytes on CytoView MEA 48-well plates. Two embryonic lines (H1 and H9) and two iPSC lines (1C and F016) were used. Spontaneous activity was recorded each day using the AxIS Navigator. (B) Spontaneous cardiomyocyte activity was first detected on Day 14 of the STEMdiff™ Cardiomyocyte Differentiation Kit protocol, and recorded up to day 25. 83 - 100% of the wells contain regular beating cardiomyocytes with very consistent excitability profiles between replicate wells by Day 25. Variability across hPSC-CM lines was small with beat periods of 0.795-1.27 sec, FPDs of 177-326 ms, and less than 10% beat period irregularity.
In summary, the Maestro MEA System and STEMdiff™ Cardiomyocyte Differentiation and Maintenance Kits could support medium to high throughput drug discovery and cardiotoxicity screening using genetically diverse hPSC-derived cardiomyocytes in a label-free manner. Data courtesy of STEMCELL Technologies, taken from Macri et al. 2017 presented at ISSCR 2017, and Recreating irregular heart beats with patient cells and gene editing.
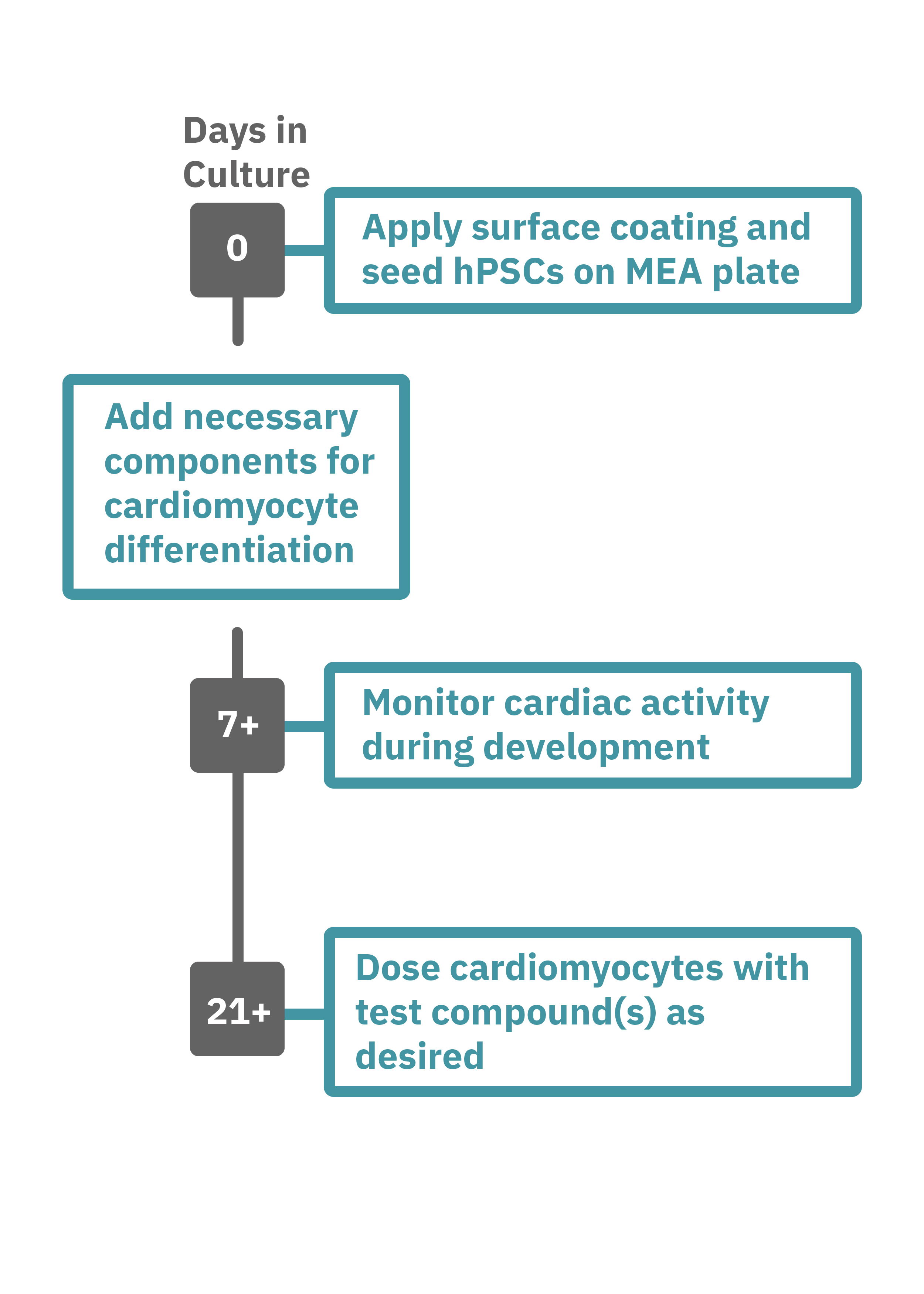
Getting started with Maestro Pro and Edge couldn't be easier. Culture your hPSCs in an Axion multiwell MEA plate (Day 0). Induce cardiomyocyte differentiation. Record as desired. Add test compounds as desired. Analyze the cardiomyocyte activity with AxIS Navigator Cardiac Module software.






