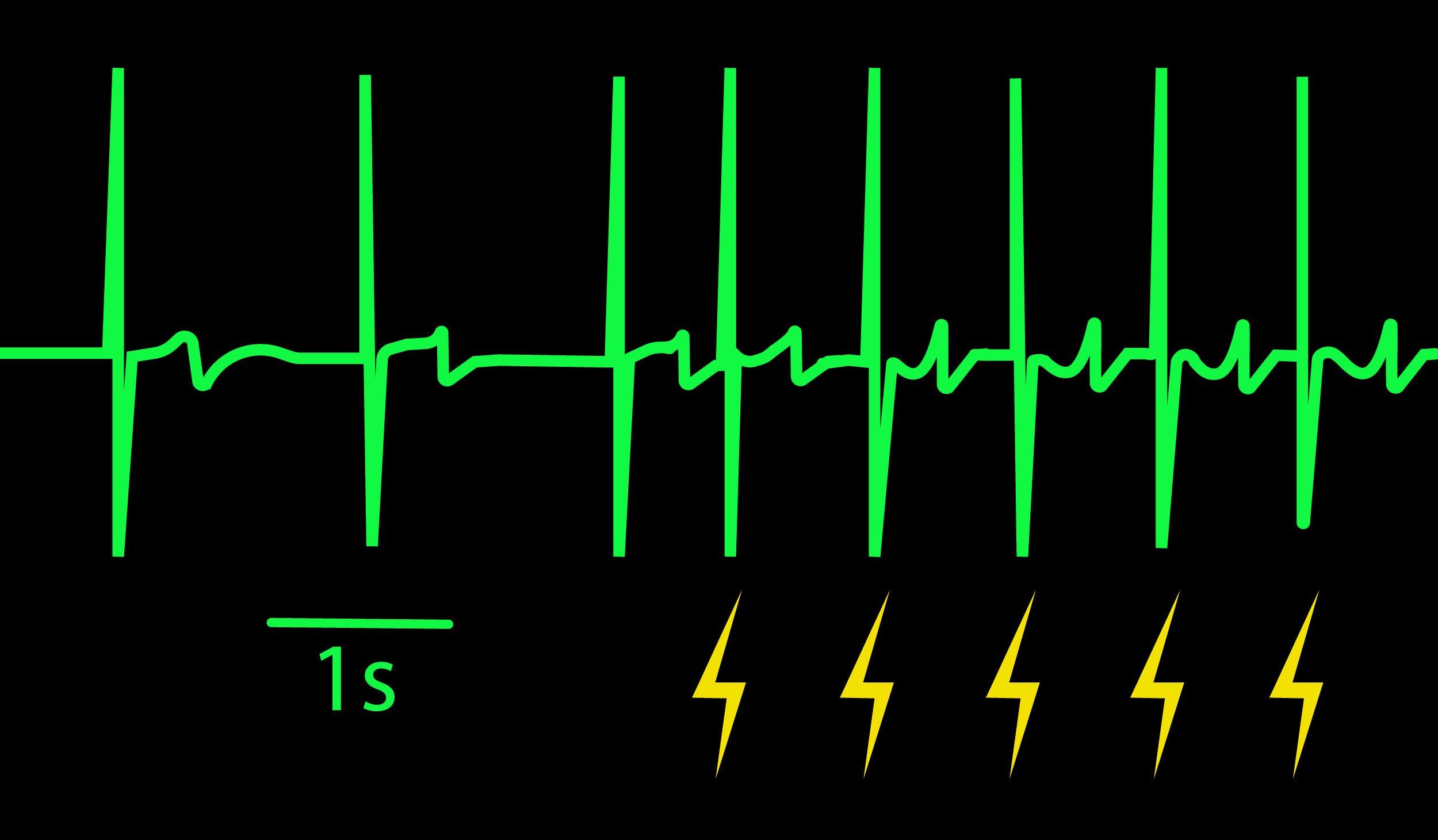
Controlling beat rate with electrical stimulation allows you to increase physiological relevance and reduce well-to-well variability. The ability to vary beat rate enables detection of use-dependent drug effects (i.e. beat rate dependent). Optogenetics can further enhance cardiac assays by providing artifact-free, precise, and targeted stimulation.
The Maestro MEA platform with the Lumos optical stimulator offer unprecedented control of cardiac beat rate with electrical or optical stimulation capabilities. Use individually or combine to precisely control cardiomyocyte beating and reduce assay variability.
-
Improve the quality and consistency of your assay>
-
Discover more by controlling cardiac activity>
-
Pacing reveals cell-specific FPD and BP relationship>
-
Quantify arrhythmic risk using cardiac pacing>
-
Assay Steps>
Compound-induced changes to cardiomyocyte field potential duration (FPD) can be used to assess the cardiac safety profile of new drugs. However, repolarization timing is tightly coupled to beat rate, meaning beat rate variability can confound FPD data interpretation. The ability to pace cardiomyocyte activity reduces both FPD well-to-well and inter-assay variability.
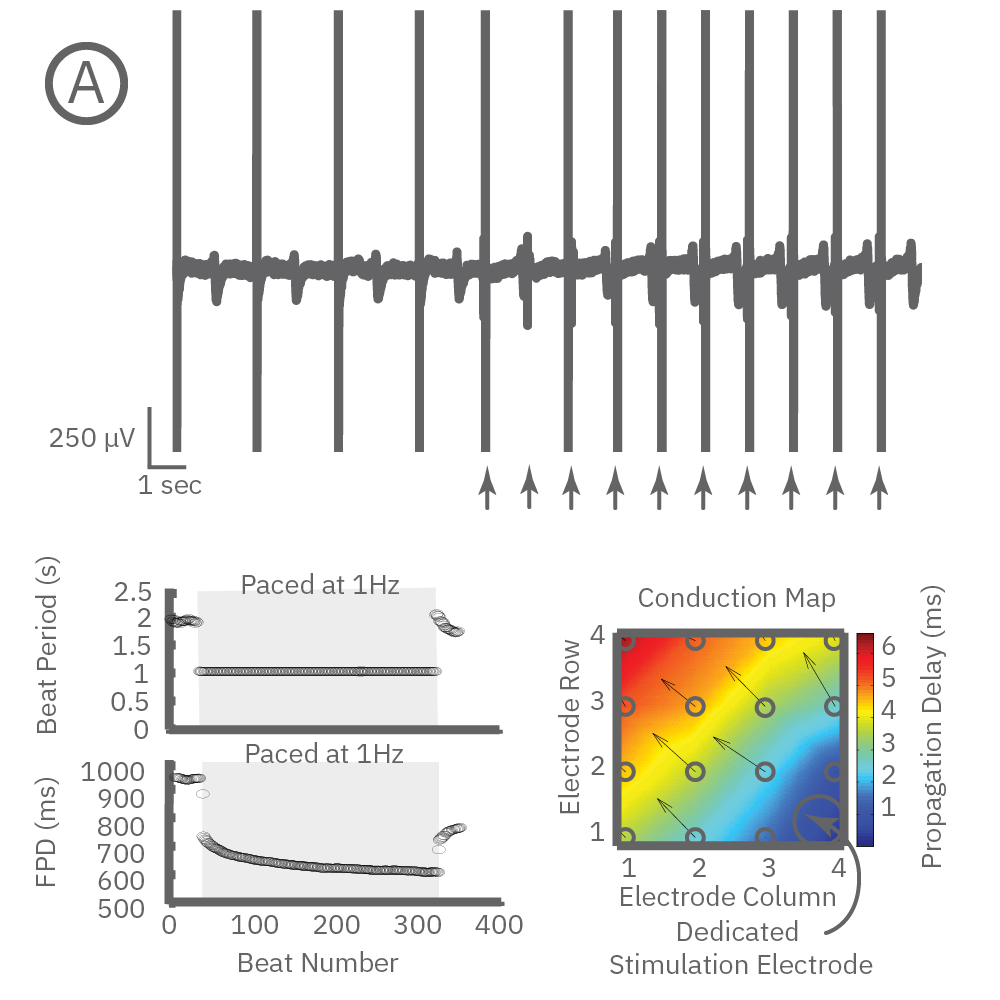
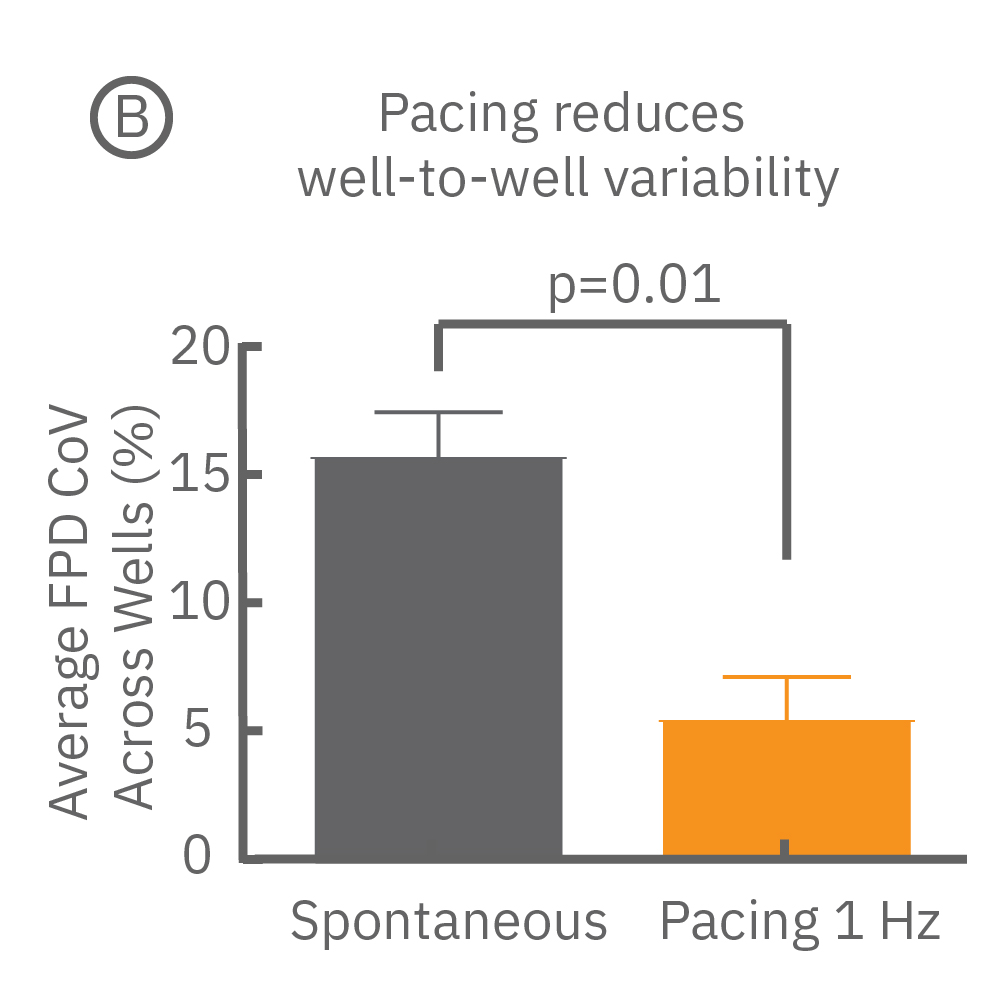
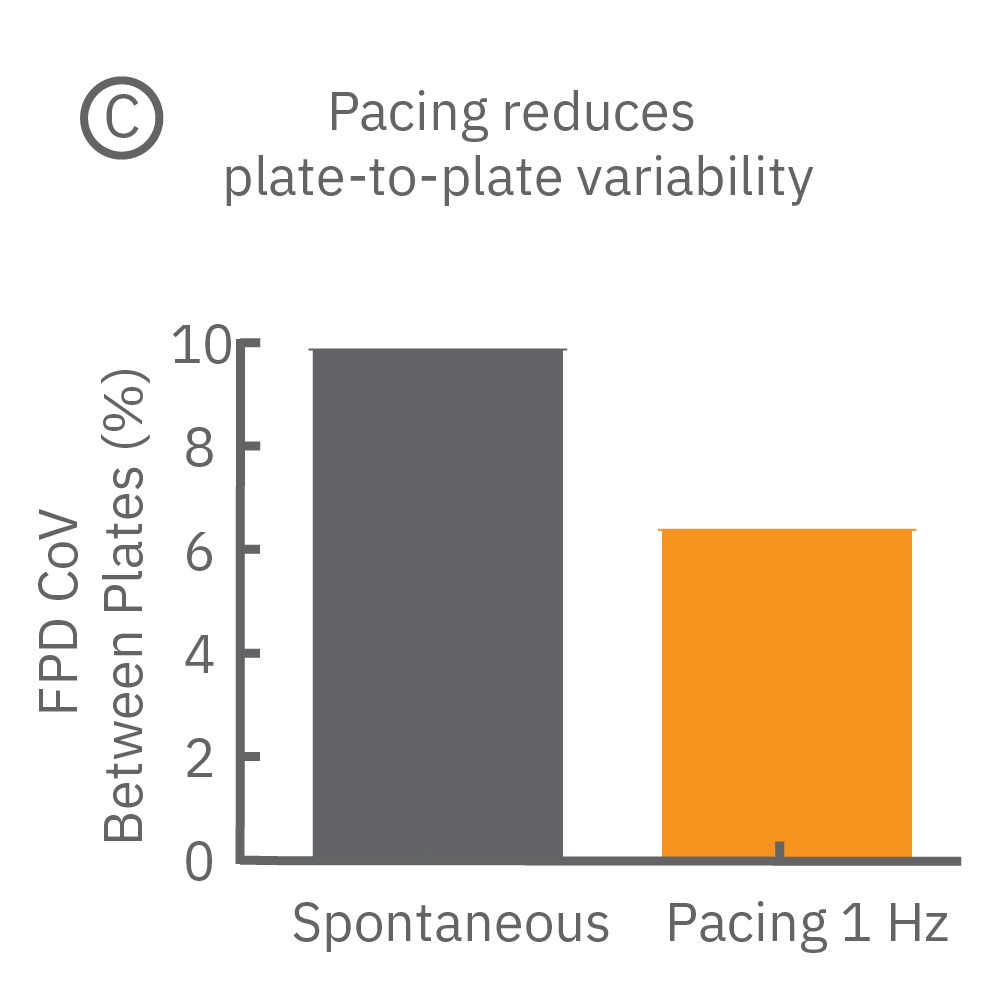
Beat period and field potential duration quickly adapt when electrical (or optical) pacing is introduced (A, arrows indicate 1 Hz pace; bottom left). (A bottom left) Key parameters, such as conduction, with real-time visualization. (B) Pacing Pluricyte® Cardiomyocytes with the dedicated E-Stim+ electrode, available in BioCircuit MEA and CytoView MEA plates, significantly reduces FPD variability between wells and plates (p=0.01) as well as variability between plates. (C) Variability was measured as the coefficient of variation (CoV) across wells and across plates.
Many compounds exhibit use-dependent effects. Reverse use-dependence, which occurs when a compound produces greater effects at slower beat rates, is an important indicator of proarrhythmic risk. The dedicated E-Stim+ electrode allows the user to systematically vary beat rate to identify such use dependent effects. Thus, pacing with the E-Stim+ electrode significantly increases the information content of your assay.
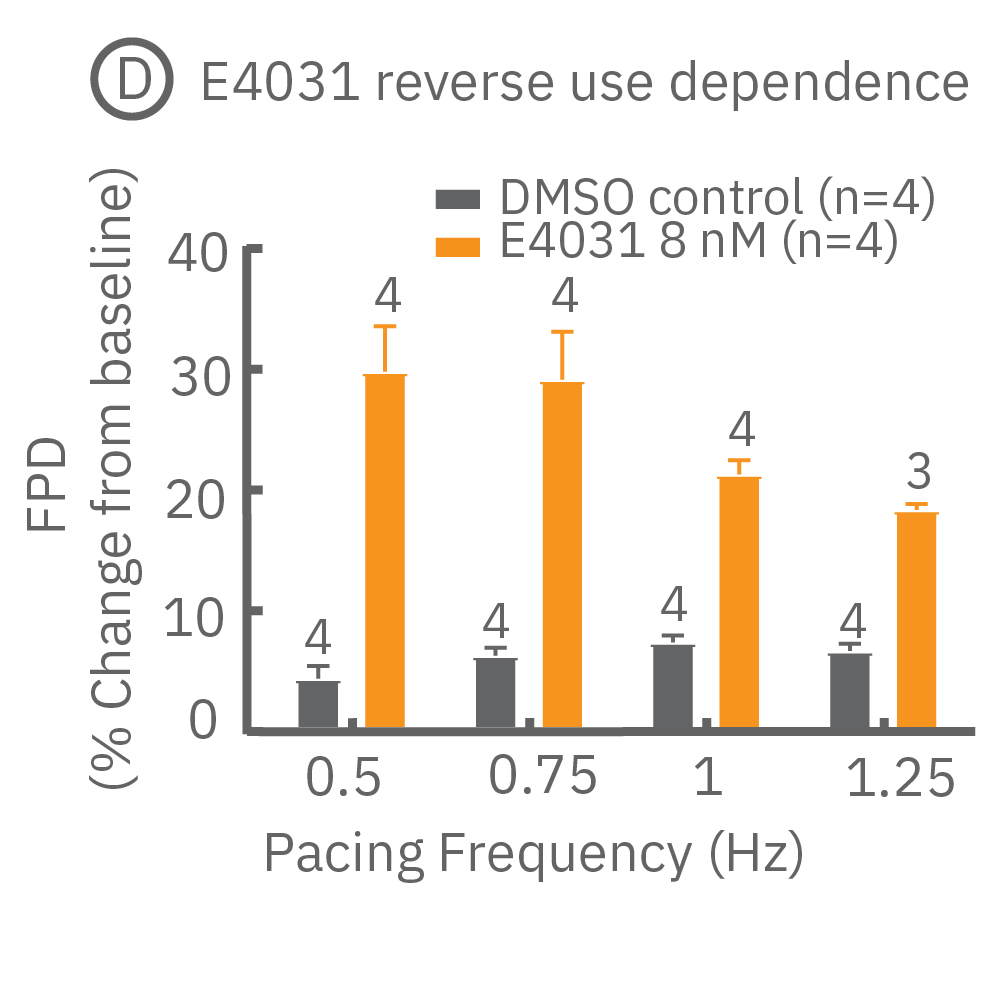
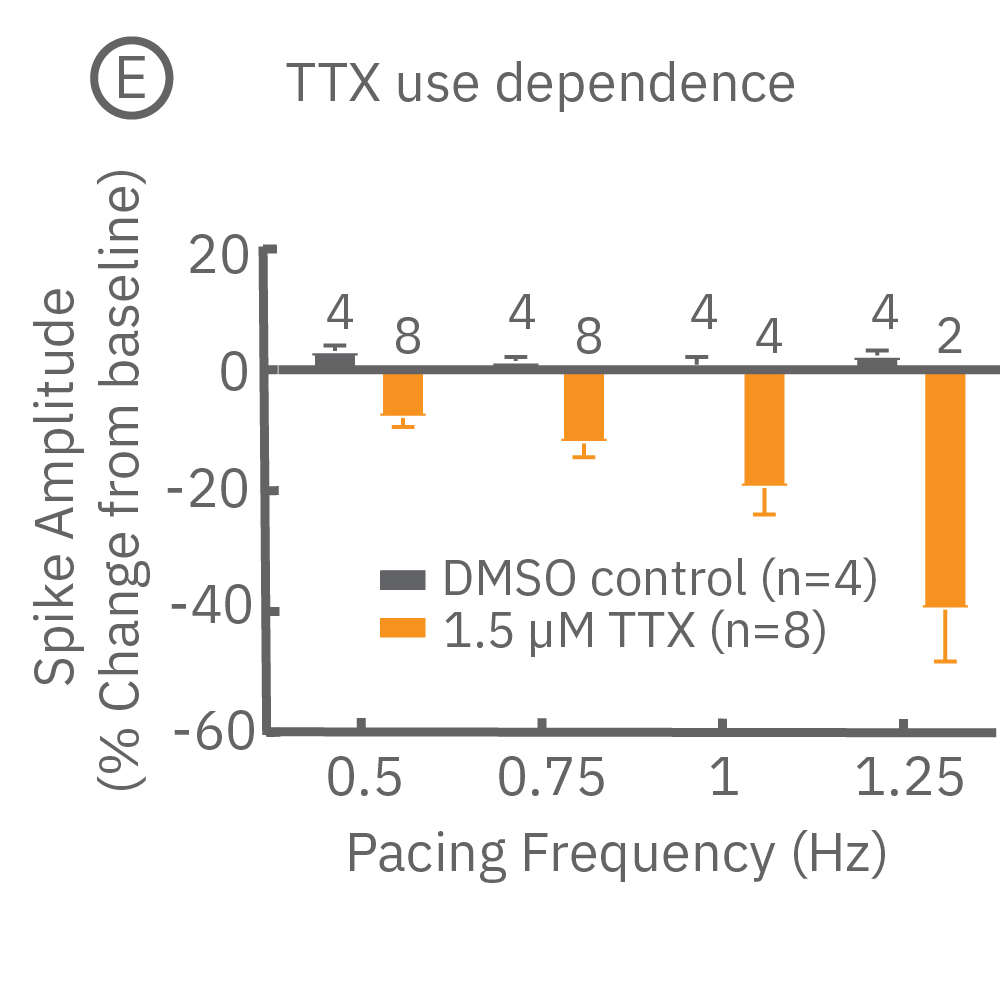
(D) Reverse use-dependence of E-4031, a hERG channel blocker, revealed by pacing Pluricyte® Cardiomyocytes with the E-Stim+ electrode. (E) Use dependence of TTX, a sodium channel blocker, revealed by pacing Pluricyte Cardiomyocytes with the the E-Stim+ electrode. (D,E) Bars represent mean ± 1 standard error of the mean. Numbers above the bars indicate the number of wells successfully paced.
Cardiac repolarization is intrinsically linked to the beat frequency, both of which are sensitive to pharmacological manipulation. Optogenetic stimulation can be used to control the beat frequency and remove it as a variable, resulting in increased reliability of the repolarization measurement.
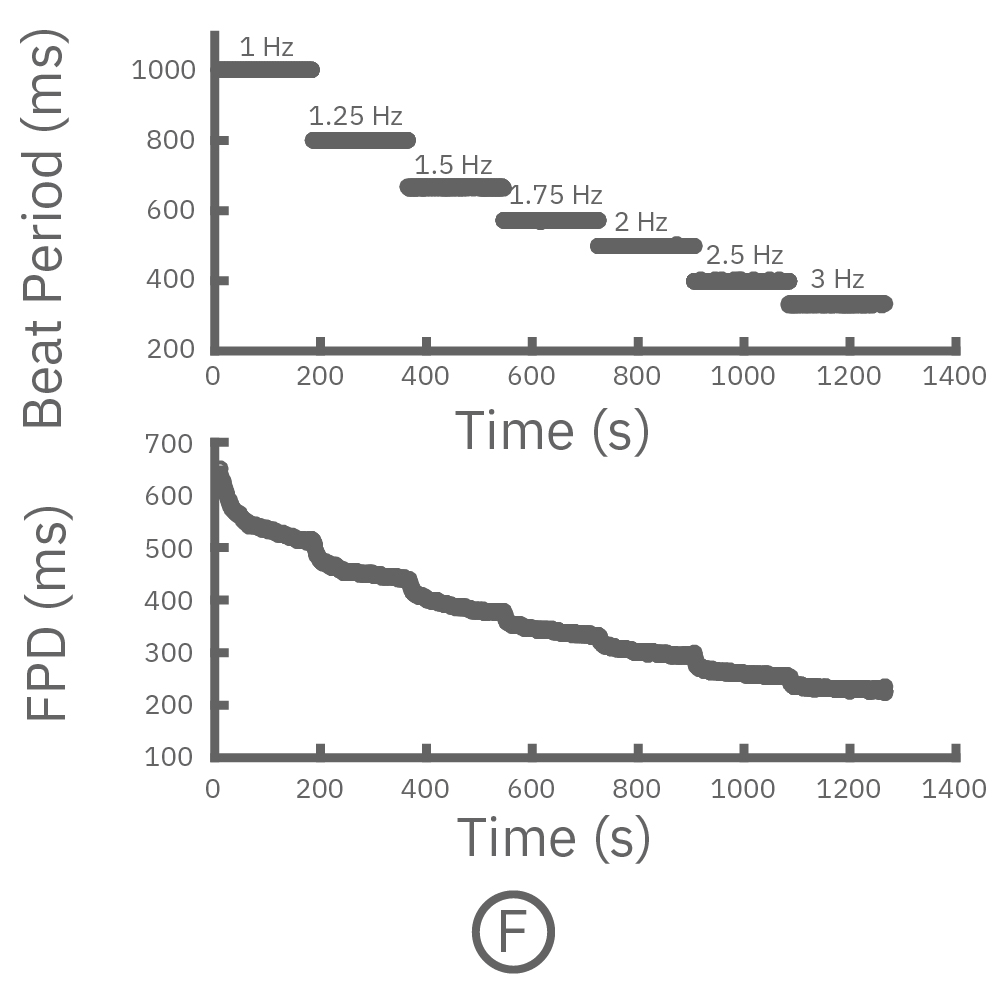
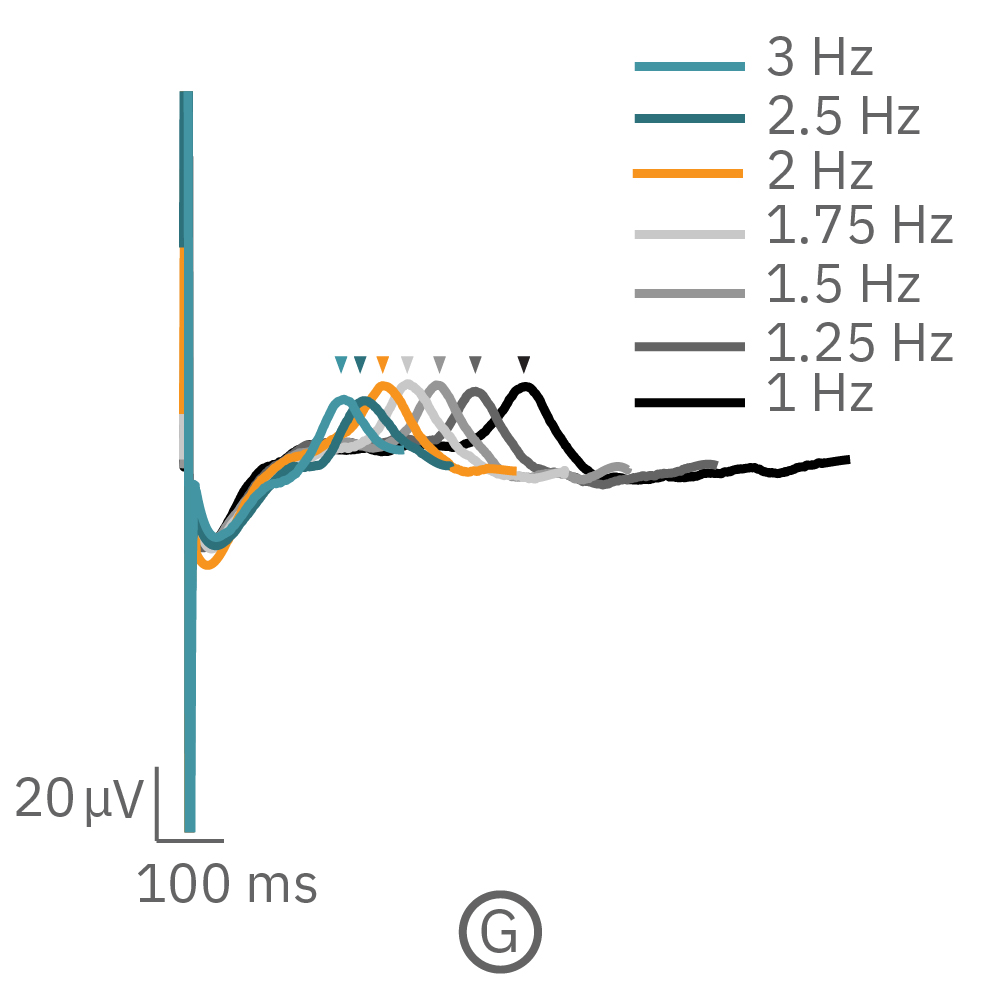
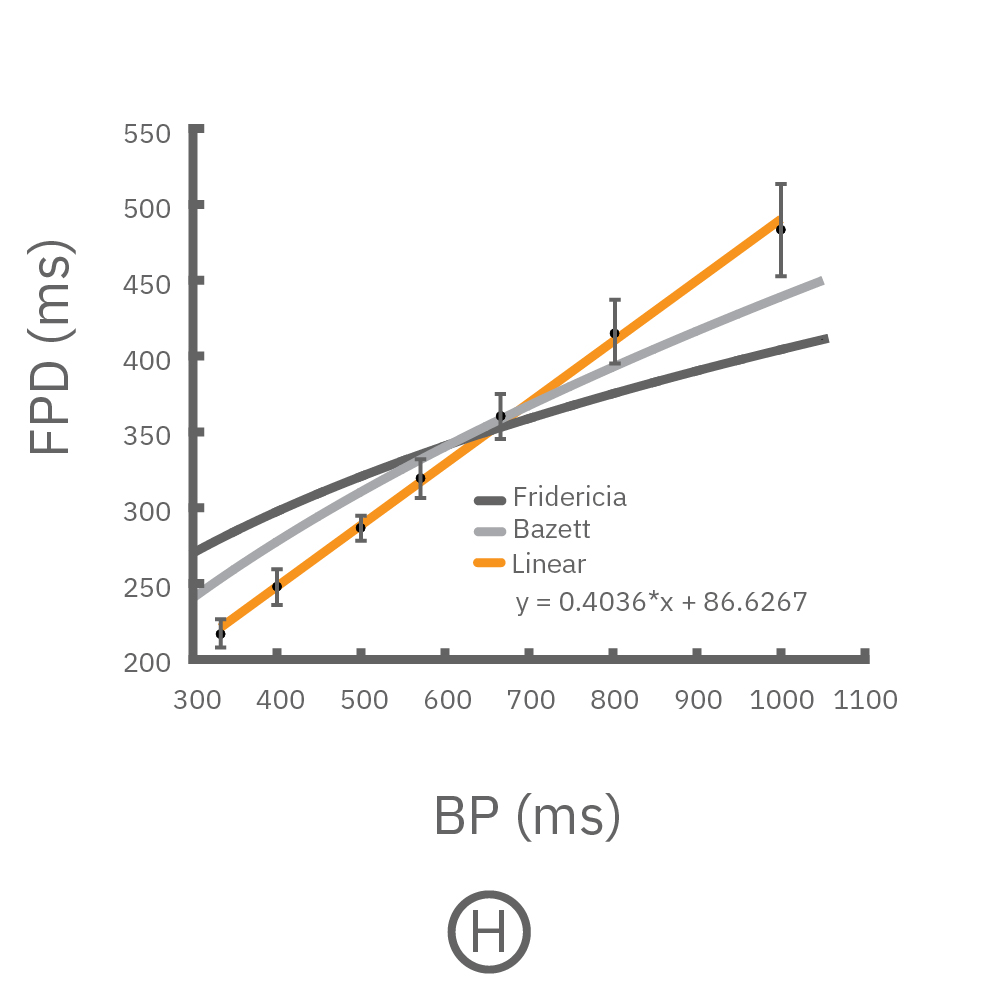
(F, G) The beat rate of Pluricyte® Cardiomyocytes was increased in a step-wise manner (known as a “chirp” assay). The field potential duration (FPD) adapted with each sequential beat rate increase up to 3 Hz. (H) Typical clinical correction formulas, the Fridericia and Bazzett, did not accurately predict the FPD. However, pacing with the Lumos revealed the cell-specific beat rate correction relationship.
Optogenetic stimulation in conjunction with LEAP can be used to quantify prolongation independent of beat period and quantify rate-dependent drug effects.
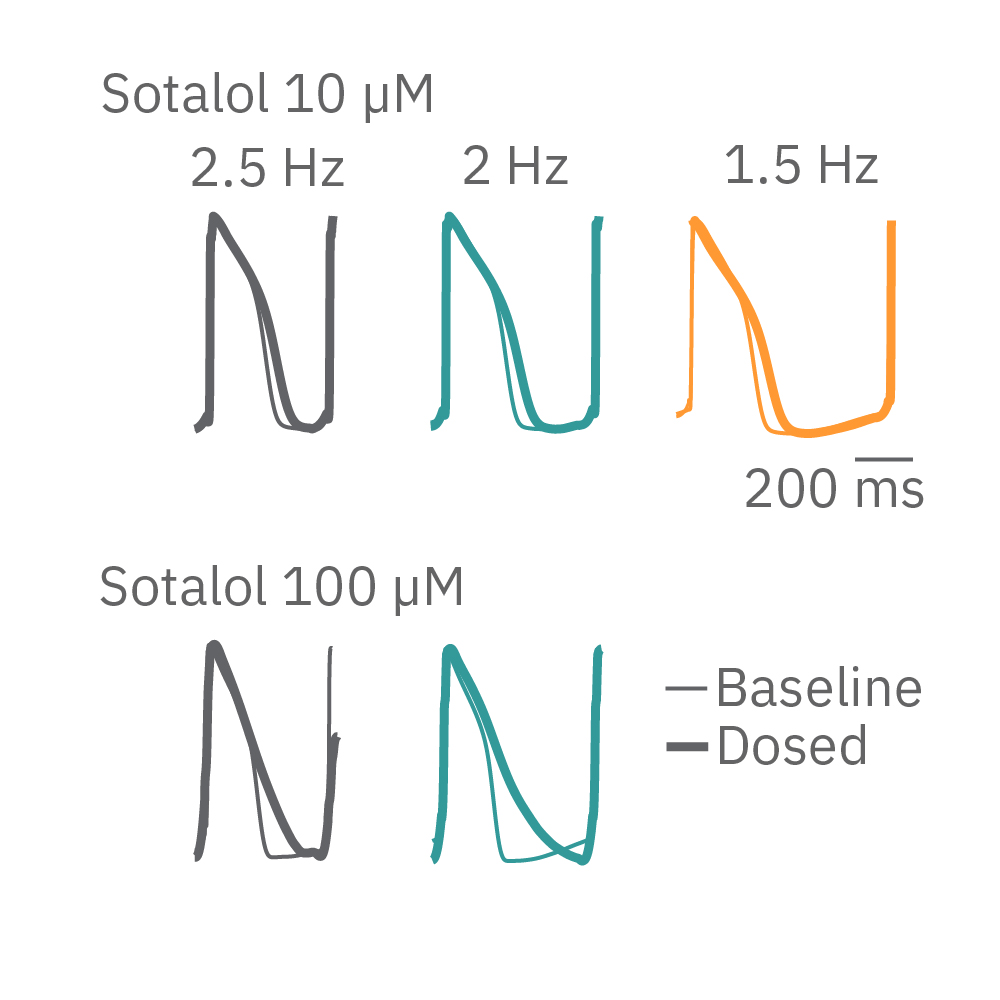
HiPSC-cardiomyocytes were transduced with channel rhodopsin 2 (ChR2), and pulses of blue light from the Lumos were used to pace cardiomyocytes at successively faster rates. Here, sotalol causes reverse use-dependent prolongation, resulting in longer prolongation at a slower beat rate.
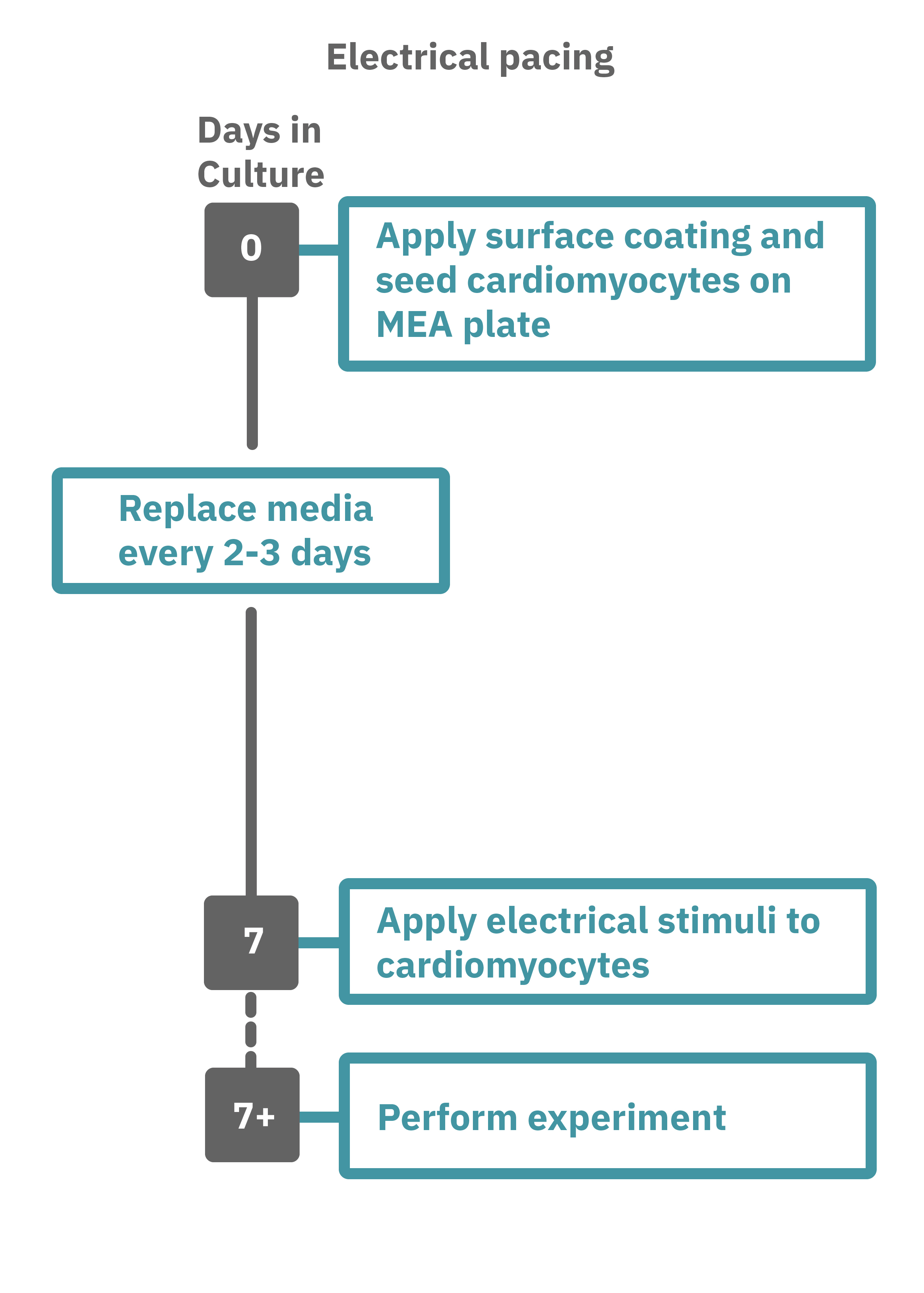
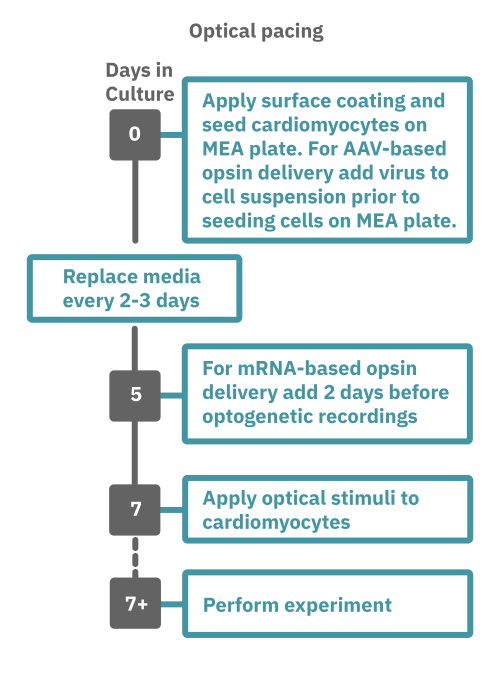
Getting started with Maestro Pro and Edge couldn't be easier. Culture your cardiomyocytes in an Axion multiwell MEA plate (Day 0). For optogenetic experiments, using adeno-associated virus-based vectors, add the virus to the cell suspension at the time of plating. Allow 7-14 days for opsin expression. When transfecting cardiomyocytes with mRNA-based opsins, transfect at peak cardiomyocyte activity and allow 2 days for opsin expression. Load the MEA plate into the Maestro MEA system at the desired recording times and begin recording. Perform optical stimulation experiments using the Lumos optical stimulator, or electrical stimulation studies using any of the microelectrodes in the wells of the MEA plate (Day 7+). Analyze the cardiomyocyte activity with AxIS Navigator Cardiac Module software.








